Pangeo is a worldwide community-centric, open, collaborative and scalable big geoscience data analytic ecosystem where scientists, developers, and research software engineers can contribute to developing software and infrastructure and work on Big Data Geoscience research problems.
Pangeo, an Open Science Gateway for Big Data science

About
Pangeo is organised as an open source software project. Check its governance documentation.
One of the main goals of Pangeo is to develop open source software and related technology for analysing large scientific datasets. Pangeo endeavours to extend the broader scientific software ecosystem and empowers everyone to access, analyse, and visualise big geoscience data. Equitable access to big data geoscience is crucial for the Pangeo community. The software developed by Pangeo is released under the BSD (or similar) open source licence, developed openly and hosted in public GitHub repositories under the Pangeo-data GitHub organisation. Some of the products released by the Pangeo community include interconnected software packages and deployments of this software stack in cloud and high-performance-computing environments and Docker container images frequently updated. Such a deployment is sometimes referred to as a Pangeo Environment.
The Pangeo ecosystem was initially developed for Geoscience. Still, it is increasingly used in different contexts (for instance, bioimaging). It has considerable potential to become a “reference” open science gateway able to leverage various infrastructures and data providers for various scientific applications.
The Pangeo community in Europe, sometimes referred to as “Pangeo@Europe”, aims to promote Pangeo in European time zones and increase the visibility of European contributors to the Pangeo ecosystem, independently from their institutions.
Another important goal of the Pangeo@Europe community is to have a shared deployment where scientists and/or technologists can exchange know-how and provide feedback. Such a Pangeo Environment can speed up the learning process because users can learn with real-world examples to access, analyse, visualise and share data, Jupyter Notebooks and best practices for making their research work FAIR and reusable. In short, it can help researchers to practise Open Science.

The challenge
Public Pangeo platform deployments that are providing fast access to large amounts of data and compute resources were all USA-based, and members of the Pangeo community in Europe did not have a shared deployment where scientists, software developers, research software engineers and/or anyone interested in big data could learn, share and exchange knowledge and best practices for delivering efficient Pangeo deployments and for practising FAIR data and software, and Open Science with big data. Before coming to EGI-ACE, there was no public documentation on how to deploy Pangeo platforms on a public cloud: documentation was limited to private cloud providers (AWS/Google), while most scientists in Europe have free access in their institutions or at national and European levels to public cloud compute and storage.
In practice, researchers had either to pay (from their research grants or themselves) to use Pangeo deployments or try (and waste time) to deploy such infrastructure themselves (but this approach is out of reach for most researchers).
The main objective of the Pangeo use case is to demonstrate how to deploy and use Pangeo on EOSC and underline the benefits for the European community.
The Pangeo use case has two main computing objectives:
- Creating a common platform for Pangeo European users;
- On-boarding European researchers on this Pangeo EOSC infrastructure.
The Solution
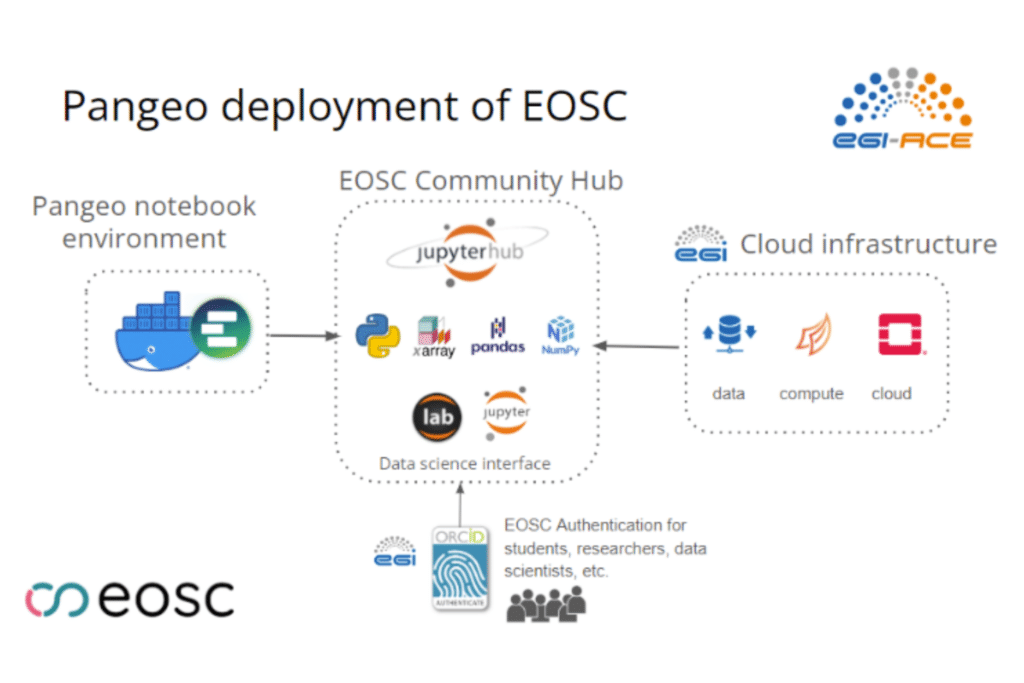
In this section, we briefly describe the Pangeo Europe Community, together with EGI, deployed DaskHub, composed of Dask Gateway and Jupyterhub, with Kubernetes cluster backend on EOSC using the infrastructure of the EGI Federation. We also used EGI Check-In to enable user registration, authenticated and authorised access to the Pangeo portal and the underlying distributed compute infrastructure.
The Pangeo EOSC Jupyterhub was deployed through the Infrastructure Manager (IM) Dashboard to enable future Pangeo deployments to be easily deployed on top of a wide range of cloud providers (AWS, Google Cloud, Microsoft Azure, EGI Cloud Computing, OpenNebula, OpenStack, and more).
The current Pangeo EOSC infrastructure has been deployed on a Kubernetes cluster on top of OpenStack. The deployment of a Kubernetes cluster and the installation of Daskhub helm chart has been done through the IM dashboard. The figure below shows the current deployment; more information is available here.
Services Provided by EGI
Login with your own credentials
Run virtual machines on-demand with complete control over computing resources
Run Docker containers in a lightweight virtualised environment
Use cloud orchestrator to deploy and configure complex virtual infrastructures
Dedicated computing and storage for training and education
Impact
consumed in the last year (March 2022-April 2023)
trained to use Pangeo until today
Pangeo Highlights
The newly deployed infrastructure has been successfully used for onboarding users during a workshop provided at the FOSS4G conference. The training material is a Jupyter Book that contains fully reproducible Jupyter notebooks about accessing data (local and remote), analyse data with Xarray, visualise data with Hvplot interactive visualisation package and understand how to scale computation with Dask. The training material is open source (CC-BY-4 licence) and has been collaboratively developed.
The FOSS4G Pangeo 101 workshop was held on Tuesday 23rd August 2022 from 14:00 – 18:00 (Florence, Italy). The course raised several interesting discussions and an increasing interest in using such infrastructure for analysing data.
The first workshop at FOSS4G gave us some visibility, for instance, among our Pangeo US colleagues: when Pangeo@US was reached to organise and deliver training to the CLIVAR community, they naturally reached out to Pangeo@Europe.
The idea of the Arctic processes in CMIP6 Bootcamp was different from a “classical” training workshop: the goal here was to have Early Career Researchers focus on “grand challenges” and to try to move the science forward using the state-of-the-art tools, and get some practical experience within their small groups as well as through the guidance of senior scientists mentors. The Pangeo community and EGI-ACE provided the technical infrastructure (Jupyterhub with Pangeo deployment similar to what was delivered for FOSS4G workshop) and technical support (best software practices for writing shareable data analysis codes and data).
The training material was adapted from the previous workshop and is also open source (CC-BY-4 licence) collaboratively developed.
Early Career Researchers and their mentors learned about CMIP6 data and how to work with CMIP6 output from many models in one go in general, and via the cloud computing infrastructure (Pangeo JupyterHub) delivered by the Pangeo Community and EGI-ACE, and the rest was spent doing actual science, i.e. apply the tools to answer the research question participants had chosen. Attendees from the CMIP6 Bootcamp were split into several groups: each group worked on different scientific questions and used different datasets. A Github organisation was created (https://github.com/orgs/clivar-bootcamp2022/repositories) to host the work of each group: access rights (private/public) were left to each group. 5 groups chose to have private repositories (to be opened with MIT licence with future publications) and 2 created public repositories.
This CLIVAR bootcamp was also the first time s3 storage was introduced to read and write files on CESNET’s Swift object storage.
More information about the bootcamp, and in particular the programme can be found here.
The eScience course on tools in Climate Science for linking observations with modelling is delivered on a yearly basis but for the first time, the EOSC infrastructure was used.
The 6th course on “eScience tools in climate science: linking observations with modelling” was held in Tjärnö, Marine Laboratory, Sweden with 17 students from Sweden, Norway, Denmark, Finland, Canada, South Africa and Australia (see Figure-2). The course was open to master students and/or new PhD students: the majority of the participants were master students. For the master students from the University of Oslo (Norway) the course was part of the GEO4990 course (10 ECTS with 5 ECTS for the eScience practicals). Students at Stockholm University (Sweden) can take this course as part of a master course MI7025 (7.5 ECTS).
A JupyterHub similar to the one delivered for the CMIP6 Bootcamp was provided to students and their mentors and professors, but this time the memory made available to the JupyterHub was increased to 48GB. Students were working in small groups, assisted by 7 highly motivated and engaged assistants. Each student had to work through a small study within a science topic, proposed by the assistant. Topics dealt with were biological production in the Arctic ocean in a world with less sea ice along with changes in ocean salinity, phytoplankton blooming and DiMethyl Sulphide (DMS)emissions. Atmospheric processes studied were aerosol size distribution variability, aerosol removal by precipitation, aerosol transport along with atmospheric rivers, but also ozone changes in the stratosphere after volcanic eruptions. Engagement and cross-talk during the course was great.
After some hiccups, all students and assistants managed to login to the EOSC Pangeo JupyterHub within the first few days. Usage of the JupyterHub throughout the course was working very well and no crashes were observed. The training material served its purpose very well and the students were able to learn from example notebooks. Similarly to the CMIP6 Bootcamp, a github organisation ( https://github.com/orgs/eScience-course/repositories ) was created for students and a repository for each group was created (7 public repositories with MIT licence). Groups stored their own Jupyter notebooks and reports in a Github repository, based on a template provided by the Pangeo community for the course ( https://github.com/eScience-course/escience2022-template). Github integration into the JupyterHub was highly appreciated. Jupyter notebooks were created by each student to document the work and results, and students delivered two presentations during the course to report on progress.
Examples were added to the tutorial during the course. The discussion section in the overall course Github repository was lively and used to exchange tips and tricks. The EOSC Pangeo s3 storage was unfortunately too complicated to set up and use. Only 2 assistants managed to login in, probably also because of limited time to test this further. Additional data was assembled and provided as read-only at the MET.NO s3 store with external help, which was more convenient for the students and assistants. However, reading data from this s3 store became slow when all the students started to analyse and load large data sets in the second week of the course. Students certainly got a glimpse of problems and solutions when dealing with large datasets. This part could probably be taught and instructed a bit better. Altogether all students finalised a report draft, participated in an internal review process and presented their findings in a hybrid meeting two weeks after the course.
More detailed information on the course can be found here and the training material (CC-BY-4) is available at https://pangeo-data.github.io/escience-2022/intro.html.
Lessons Learned and Next Steps
The collaboration with EGI-ACE is a real success for Pangeo. It delivered efficient and scalable Pangeo JupyterHubs that were successfully used to onboard more than 100 users from more than 10 countries. The feedback from users is overall very positive. During the FOSS4G, several attendees struggled with EGI Check-in; mostly because it takes some time between the registration and effective access to the resource. For later events, we anticipated and asked attendees to register to the EOSC services at least one week (ideally 2 weeks before the workshop) before the start of the training/event to troubleshoot any potential issues. In addition, we improved the documentation on how to join EOSC Pangeo JupyterHub and connect with EGI Check-in.
Access to data was reported to be very difficult and/or inefficient e.g. the solution offered by EGI-ACE was not meeting users’ expectations. This is why the Pangeo community submitted a use case to the C-SCALE project e.g. to get data close to the compute resources and overall provide users with easier and faster access to data.
More details on the lessons learned will be provided at EGU 2023 (Fouilloux, A., Marasco, P. L., Odaka, T., Mottram, R., Zieger, P., Schulz, M., Coca-Castro, A., Iaquinta, J., and Eynard Bontemps, G.: Pangeo framework for training: experience with FOSS4G, the CLIVAR bootcamp and the eScience course, EGU General Assembly 2023, Vienna, Austria, 24–28 Apr 2023, EGU23-8756, https://doi.org/10.5194/egusphere-egu23-8756, 2023)
In parallel to onboarding events, the Pangeo community continues its collaboration with EGI to improve the Pangeo deployment and facilitate Open Science practices with, for instance, the deployment of a Binder instance with a Dask gateway and provide a common approach to spatial data analysis, independently of data and infrastructure providers.

With support from the EGI-ACE, the project Pangeo@Europe provides scientists with big-data analysis and ready-to-use Pangeo deployments on the European Open Science Cloud (EOSC).
These deployments come with a DaskHub, including Dask Gateway and JupyterHub, with a Kubernetes cluster backend using a comfortable amount of compute and storage resources from the EGI Federation.
The Pangeo software stack, which was initially put together to fulfil users' needs in the geoscience community, is so comprehensive that it now attracts interest from other scientific disciplines (DNA sequencing, bioimaging analysis, physics, etc.).
News from Pangeo
Loading